Challenge Open Access
Robust Detection of Heart Beats in Multimodal Data: The PhysioNet/Computing in Cardiology Challenge 2014
George Moody , Benjamin Moody , Ikaro Silva
Published: Jan. 7, 2014. Version: 1.0.0
Final results for the 2014 PhysioNet/CinC Challenge (April 15, 2015, 2:28 p.m.)
The top results of the follow-up entries as well as all entries from phases I-III (at the end of February 2015) were achieved by Urska Pangerc (93.64), Alistair Johnson (91.50), Sachin Vernekar (90.97), Christoph Hoog Antink (90.70), and Abid Rahman (90.16). The complete updated rankings are now available including follow-up entries as well as all entries from phases I-III.
Official results for Phase III of the 2014 PhysioNet/CinC Challenge (Sept. 9, 2014, 2:31 p.m.)
The top official results in Phase III were achieved by Alistair Johnson (87.9), Teo Soo-Kng (86.7), Thomas De Cooman (86.6), Jan Gierałtowski (86.4), Marcus Vollmer (86.2).
More news
Preparing your talk for Computing in Cardiology 2014 (Aug. 21, 2014, 2:26 p.m.)
Information on how to prepare your talk for Computing in Cardiology is described here.
Official results for Phase II of the 2014 PhysioNet/CinC Challenge (June 25, 2014, 2:22 p.m.)
The top official results in Phase II were achieved by Thomas De Cooman (86.2), Marcus Vollmer (86.1), Urska Pangerc (85.2), Filip Plesinger (85.0), and Alistair Johnson (84.6).
Official results for Phase I of the 2014 PhysioNet/CinC Challenge (April 17, 2014, 2:19 p.m.)
The top official results in Phase I were achieved by Marcus Vollmer (93.2), Urska Pangerc (89.2), Lars Johannesen (88.9), Quan Ding (88.9), and Teo Soo-Kng (88.7). Several other high-scoring entries were disqualified for reasons described in the FAQs.
Please include the standard citation for PhysioNet:
(show more options)
Goldberger, A., Amaral, L., Glass, L., Hausdorff, J., Ivanov, P. C., Mark, R., ... & Stanley, H. E. (2000). PhysioBank, PhysioToolkit, and PhysioNet: Components of a new research resource for complex physiologic signals. Circulation [Online]. 101 (23), pp. e215–e220.
Introduction
This challenge aims to encourage the exploration of robust methods for locating heart beats in continuous long-term data from bedside monitors and similar devices that record not only ECG but usually other physiologic signals as well, including pulsatile signals that directly reflect cardiac activity, and other signals that may have few or no observable markers of heart beats. Our goal is to accelerate development of open-source research tools that can reliably, efficiently, and automatically analyze data such as that contained in the MIMIC-II Waveform Database, making use of all relevant information.
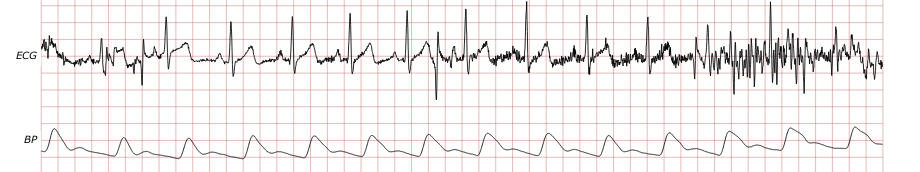
Most existing beat detectors are QRS detectors, operating on the ECG. In this challenge as in many clinical and research settings, although the ECG is usually available, its quality may vary considerably over time, and on occasion the ECG may be missing entirely. Thus even an excellent QRS detector is not sufficient to be a successful challenge entry, since examination of pulsatile signals such as continuous blood pressure (BP) and photoplethysmograms (PPGs) can help to fill in the gaps. Even signals that are not acquired as indicators of cardiac activity, such as EEG signals, are often contaminated by ECG components that can be detected and exploited. In most subjects, the observed relationships between respiration and heart rate can be used to model heart rate, and together with nearby context derived from ECG or other cardiac signals, these models can predict beat locations from respiratory signals.
Rules and Deadlines
All deadlines occur at noon GMT (UTC) on the dates mentioned below. If you do not know the difference between GMT and your local time, find out what it is before the deadline!
The Challenge will take place in three phases, with a week-long hiatus between phases I and II, and a shorter hiatus between phases II and III. The week-long hiatus is necessary to ensure that all Phase I entries can be scored before the CinC abstract deadline (see below); the shorter hiatus is to allow all Phase II entries to be scored before Phase III begins. Entries will not be accepted during either hiatus.
| Start at noon GMT on |
Entry limit |
End at noon GMT on |
|
| Phase I | 7 January | 5 | 7 April |
| [Hiatus] | 7 April | 0 | 16 April |
| Phase II | 16 April | 5 | 22 |
| [Hiatus] | 22 |
0 | 23 |
| Phase III | 23 |
5 | 15 August (final deadline) |
All entries must be received no later than the end of Phase III (noon GMT on Friday, 15 August 2014). In the interest of fairness to all participants, late entries will not be accepted or scored.
Each participant may receive scores for up to five entries submitted during each of phases I, II, and III. Unused entries may not be carried over to later phases. Entries that cannot be scored (because of missing components, improper formatting, or excessive run time) are not counted against the entry limits.
To encourage collaboration among participants, we will publish each participant's most successful entry, and its ranking, at the start of Phase II on 16 April and Phase III on 23 16 June. Study these entries as the Challenge continues, and learn from them; then form alliances with other participants whose ideas may complement yours.
The authors of the top entries at the ends of Phases I, II, and III will receive awards of up to US$1000 during the closing ceremony of Computing in Cardiology 2014, on Wednesday afternoon, 10 September 2014, in Cambridge, Massachusetts.
To be eligible for an award, you must do all of the following:
- Submit at least one entry that can be scored before the phase I deadline (noon GMT on Monday, 7 April 2014).
- Submit an acceptable abstract (about 300 words) on your work on the Challenge to Computing in Cardiology no later than 14 April 2014. Include the overall score for at least one phase I entry in your abstract. Please select "PhysioNet/CinC Challenge" as the topic of your abstract, so it can be identified easily by the abstract review committee. You will be notified if your abstract has been accepted by email from CinC during the first week in June.
- Submit a full (4-page) paper on your work on the Challenge to CinC no later than 1 September 2014.
- Attend CinC 2014 (7-10 September 2014) and present your work there.
For each of Phases I and II, we will award US$300 to the first-place team, $200 to the second-place team, and $100 to each of the third-, fourth-, and fifth-place teams. For Phase III, these awards will be $400, $300, and $200 respectively. Eligible teams may receive at most one award for each phase.
Please note that the CinC abstract deadline is more than two weeks earlier in 2014 than its traditional date!
Challenge Data
Data for this Challenge are 10-minute (or occasionally shorter) excerpts ("records") of longer multiparameter recordings of human adults, including patients with a wide range of problems as well as healthy volunteers. Each record contains four to eight signals; the first is an ECG signal in each case, but the others are a variety of simultaneously recorded physiologic signals that may be useful for robust beat detection. Signals have been digitized at rates between 120 and 1000 samples per second; in any given record, however, all signals are sampled at the same, fixed frequency.
A training data set for this challenge is available for study. It is a set of 100 records, named 100, 101, ..., 199, and it is provided in the "Files" section below. You may wish to explore these records visually using LightWAVE. This data set is also available for download as a zip archive and as a tarball (about 111 MB in either format).
A new augmented training set, consisting of 100 records from the original \ test set has been released.The annotations were not generated from any specific channel and there was no fixed fiducial point, since some of the annotations were placed manually. The annotations include only beat labels and do not differentiate between beat types (all annotated beats were arbitrarily set to normal, 'N' beats).
The training set includes many records that can be processed without errors by the sample entry using the ECG only, but others will pose serious difficulty unless your entry makes good use of available information in the other signals; a few of the difficult records are 112, 133, 169, and 188.
A set of reference beat annotations for the training set is also available. In this Challenge, reference beat annotations represent the preponderance of expert opinions about the locations of the observed (or imputed) QRS complexes in the ECG signal.
A separate hidden test data set has been assembled for evaluating Challenge entries. Performance of the challenge entries on this hidden test set determines their rankings and thus the winners of the Challenge. The test set will not be available for study by participants, in order to avoid the possibility that entries will be "tuned" (optimized for high performance) on the test data and that the results will therefore be less predictive of performance on unknown data. Previously unused records will be added to the hidden data set at the start of Phases II and III.
Important differences between the training set and the test set: The training set is intended to give participants an opportunity to see some of the problems their entries will face in the challenge, and to give us a way to verify that submitted entries are working as their authors intend. The performance of challenge entries on the training set does not contribute in any way to their scores and ranks in the Challenge.
The test set contains a wider variety of signals than in the training set. A successful entry needs to be able to discover their relationships and exploit features that can predict beat locations. Unlike the training set (sampled at a uniform 250 samples per second per signal), signals in the test set have been sampled at rates between 120 and 1000 samples per second.
Entering the Challenge
To participate in the challenge, you will need to create software that is able to read the test data and record the times of occurrence of the beats in PhysioBank-compatible annotation files, without user interaction, in our test environment. A sample entry (entry.tar.gz or entry.zip) that demonstrates how this can be done is available to help you get started. It must be possible to run your software on GNU/Linux using open-source components only.
It is not necessary to develop your entry using a Linux platform. The sample entry runs on FreeBSD, Mac OS X, and MS-Windows as well as GNU/Linux. If you avoid the use of platform-specific code and libraries, your entry should be able to run in our GNU/Linux test environment.
You may use any programming language (or combination of languages) supported using open source compilers or interpreters on GNU/Linux, including C, C++, Fortran, Haskell, Java, Octave, Perl, Python, and R. Other languages may be available.
Your entry must be able to run from beginning to end without requiring user interaction. It doesn't need, and should not require, a graphical user interface.
Your entry should attempt to locate all of the beats in its input, starting from the beginning of each recording. No learning period is allowed; missing the first beat or the last one will affect your score to the same extent as missing any other beat. Locating every beat, even at the beginning of a recording, is difficult in real-time clinical applications. For off-line analysis of recorded data for research as well as some clinical applications, however, there is no need for real-time analysis per se. In this challenge, there are no real-time constraints or limitations with respect to decision delay. Your code may read any portion of the input record, even the entire record, before recording any beat locations. Its analysis can be sequential or non-sequential, forwards, backwards, etc.
Your entry's annotations do not need to be placed at the precise locations of the reference beat annotations; in order to be counted as correct, they may be located anywhere within 150 ms of a reference annotation. Note, however, that all of the reference annotations are aligned with respect to the observed or expected QRS locations; thus, for example, if a blood pressure (BP) waveform was used to locate a beat that could not be observed directly in the ECG because of noise or signal loss, the reference annotation for that beat would be located at the expected position of the QRS complex, based on nearby measurements of the delay between the QRS complex and the BP waveform feature used for beat detection. Since this delay can easily exceed 150 ms, your entry needs to make similar adjustments to beat locations when it relies on non-ECG signals.
The output annotation files must contain annotations in time order, but you don't need to worry about this point unless your entry attempts to generate annotation files without using the WFDB library. The WFDB library's wfdbquit function (which should be invoked to close open files) sorts annotations automatically so that they are written in time order in the annotation file.
To handle signals sampled at varying frequencies, you may use the WFDB library's sampfreq function to determine the sampling frequency of the input signals, and then adjust any parameters that depend on the sampling frequency accordingly; see gqrs.c in the sample entry for an example. Alternatively, you can use the WFDB library's setsampfreq function to force the input data to be resampled at your preferred sampling frequency; see wabp.c for an example of this approach. The first approach is preferred, since it does not introduce resampling noise, and it will be faster than the second, but the differences are likely to be minor.
Preparing an entry for the challenge
When you submit your entry, it must include your software in source form, as well as a complete set of the 100 annotation files that you have created by running your software, using the training data set as input, and a pair of scripts (batch files) that can be used to replicate the annotation files in our test environment, given your software in source form, and the test data set. Since entries are tested and scored automatically, it is essential to submit them in a format that can be understood by our literal-minded autoscorer. Please follow the guidelines in this section very carefully!
Participants should download the sample entry (entry.tar.gz or entry.zip). Entries should mimic the layout of the sample entry; specifically, they must contain:
- setup.sh, a bash script run once before any other code from the entry; use this to compile your code as needed
- next.sh, a bash script run once per training or test record; it should analyze the record using your code, saving the results as a beat annotation file with the annotator name qrs
- AUTHORS, a plain text file listing the members of your team who contributed to your code, and their affiliations
- sources/, a directory containing your code in source form; this should include sources for any custom or non-standard libraries needed (and setup.sh should include commands for building and linking any such libraries to your code)
- challenge/2014/set-p/, a directory containing the annotation files for each record in the training set; these must be named 100.qrs, 101.qrs, ..., 199.qrs. These files are used for validation only, not for ranking entries (see below).
See the comments in the sample entry's setup.sh and next.sh to learn how to customize these scripts for your entry.
See gqrs.c for an example of how to read the data and generate annotation files in the correct format using the WFDB library functions wfdbinit, getvec, putann, and wfdbquit. The WFDB library, and simpler examples of its use, can be found in the WFDB Software Package.
In order to submit an entry written in m-code successfully, it must be able to run using GNU Octave.
A sample entry written in Octave m-code is available here. The entry illustrates how to install and use the WFDB Toolbox for MATLAB and Octave to read the Challenge data and to write beat annotation files.
We verify that your code is working as you intended, by comparing the annotation files that you submit with your entry with annotation files produced by your code running in our test environment using the same inputs. If your code passes this validation test, it is then evaluated and scored using a large hidden test data set, and these scores determine the ranking of the entries and the outcome of the Challenge.
In addition to the required components, your entry may include a file named DRYRUN. If this file is present, your entry is not evaluated using the hidden test data, and it will not be counted against your limit of five entries per phase; you will receive either a confirmation of success or a diagnostic report, but no scores. Use this feature to verify that none of the required components are missing, that your setup.sh script works in the test environment, and that your next.sh script produces the expected output for the training data within the time limits.
Submitting an entry
Please do not submit entries by email; they will not be read or scored.
All submissions must be made via the PhysioNet/CinC Challenge 2014 project on PhysioNetWorks. Make a free personal account for yourself and log in at https://physionet.org/users/, then click on the PhysioNet/CinC Challenge 2014 link in the Projects section of your PhysioNetWorks home page. Join the project at any time while the Challenge is open to create your participant page.
Upload your entry for automated scoring using the form on your participant page. Note that it must be named entry.tar.gz or entry.zip, and its contents must be laid out as described in the previous section. Improperly-formatted entries, including entries that are missing required components, will not be scored.
Test environment
Entries are evaluated in a GNU/Linux environment (64-bit Debian 7.3 "Wheezy"), with a typical set of open-source software development compilers, interpreters, libraries, plugins, and utilities (including gcc, gfortran, ghc, java, make, octave, perl, python, and R. We will consider reasonable requests to add other packages from the Debian stable repository to the test environment at any time during the first four weeks of each Challenge phase.
Additions to the test environment: armadillo and boost libraries (13 January); liboctave-dev (30 January); gsl-bin (3 February); octave-statistics (15 March); imagemagick, imagemagick-common, libdjvulibre-text, libdjvulibre21, libexiv2-12, libilmbase6, liblensfun-data, liblensfun0, liblqr-1-0:amd64, libmagickcore5:amd64, libmagickcore5-extra:amd64, libmagickwand5:amd64, libnetpbm10, libopenexr6, netpbm, octave-control, octave-general, octave-image, octave-miscellaneous, octave-optim, octave-signal, octave-specfun, octave-struct, ufraw-batch, units (22 April)
In addition to these standard Debian packages, the test environment includes the WFDB Software Package including the WFDB library. The WFDB library can be linked with C or C++ code using -lwfdb as an option to gcc, as illustrated in setup.sh in the sample entry. Your entry will not have access to a network connection.
Evaluation of your entry will be conducted by a supervisor script that looks similar to this pseudocode:
stage 1:
unpack entry.tar.gz or entry.zip, report error and exit if unable to do so
check that all required components are present, report error and exit if
any are missing
run "setup.sh", report error and exit if it fails or needs more than five
minutes
stage 2:
for each record R in the training set
run "next.sh R", report error and exit if it fails or needs more than
40 20 seconds
compare R.qrs with the copy of R.qrs from entry.tar.gz or entry.zip,
report error and exit if they are different
exit if the entry includes a file named "DRYRUN"
stage 3:
set cumulative run time to zero
for each record R in the hidden test set
run "next.sh R"; if its running time reaches 40 20 seconds, or if it
fails, interrupt it and create an empty R.qrs
update cumulative run time, report error and exit if more than 1 hour
stage 4:
for each record R in the hidden test set
compare R.qrs with the reference annotations, count TP, FN, FP
remove any files created by next.sh
calculate and report gross and average Se and +P, and overall score
exit
The supervisor script keeps track of the time needed by your setup.sh and next.sh scripts in order to avoid spending an excessive amount of time attempting to run a misbehaving entry. An error exit will result in no score for the entry, and the entry will not count against your limit of five entries in each phase of the Challenge. Most entries will not approach the time limits; in our test environment, the sample entry's setup.sh requires only a second or two, and its next.sh processes a 10-minute record in about 35 milliseconds. We will consider reasonable requests to adjust the time limits on a case-by-case basis.
Scoring
If your entry is properly formatted, and nothing is missing, it is tested and scored automatically, and you will receive your scores when the test is complete (depending on your entry's run time, this may take an hour or more). If you receive an error message instead, read it carefully and correct the problem(s) before resubmitting.
The annotations generated by your code are compared beat-by-beat with reference annotations that reflect the consensus of several expert annotators, using bxb(1) and sumstats(1) from the WFDB Software Package. As noted above, no learning period is allowed (bxb is run with the -f 0 option so that comparison begins with the first annotation in each record).
Your entry's scores are reported as gross and average sensitivity, and gross and average positive predictivity (four numbers, each between 0 and 100; see below). For purposes of ranking, we average these four performance statistics together with equal weight to obtain an overall score. The top-ranked entry will have the highest overall score.
Beat-by-beat comparison and calculation of performance statistics
The bxb application matches your entry's annotations ("test annotations") with reference annotations to count, for each record in the test data set, the numbers of correctly-detected beats (true positives, or TP), missed beats (false negatives, or FN), and detections of non-beats (false positives, or FP). To match a reference annotation, a test annotation must be located within 150 ms of the reference annotation, and must be the nearest test annotation to the reference annotation.
In this challenge, sensitivity (Se) is the percentage of beats that are true positives, and positive predictivity (+P) is the percentage of detections that are true positives:
Gross statistics are derived from the sums of all TP, FP, and FN over the test data set; average statistics are the means of the statistics calculated individually for each record of the test data set.
If an entry fails to generate an annotation file for a given test set record, or if it does not complete its analysis of the record within the 40-second 20-second time limit, any output it has produced is replaced with an output annotation file that contains a single annotation that does not match any reference annotation. In such cases, the beat-by-beat comparison yields 1 FN for each beat marked in the reference annotation file for the record, 0 TP, and 1 FP. The Se and +P for the record are both 0 (affecting the average Se and +P).
Frequently asked questions about the Challenge
How can I determine if my entry is fast enough?
As noted above, the sample entry processes a ten-minute record in about 35 milliseconds in our test environment, or about 300 times faster than necessary on average. Measure the run time of the sample entry on your computer, and compare it to the run time of your entry on the same data. If your entry requires no more than 300 times as long to run than the sample entry on average, it is probably fast enough.
Note that the 40-second 20-second limit per record is intended to allow extra time for processing unusually complex records. If your entry's mean run time is greater than about 10 seconds per record in the test environment, it is unlikely to complete its processing of the full 300-record test set in phase III within the one-hour limit.
Can I get diagnostic output from my entry?
Yes, in stages 1 and 2 (see above), but not in stages 3 and 4. The general rule is that diagnostic output is available if it was produced without access to test set data.
In stage 1, if your setup.sh fails (exits with non-zero status), anything it writes to the standard output is reported and the evaluation stops. Otherwise, no output from setup.sh is reported.
Similarly, in stage 2, if your next.sh fails on a training set record, anything it writes to the standard output is reported and the evaluation stops (the remaining records are not tested). Otherwise, no output from next.sh is reported.
In stage 3, however, if your next.sh is unable to process a test set record within 40 20 seconds, it will receive poor partial scores for that record (see above), but the evaluation will continue.
If your entry exceeds the one-hour time limit for processing the test set in stage 3, the percentage of records completely processed within one hour is reported. Otherwise, if your entry failed to process one or more test set records, or if it did not complete its analysis of one or more records within the 40-second 20-second per-record time limit, the numbers of failures and timeouts will be reported, along with the scores. No other diagnostic output relating to test set records will be reported.
May I submit a binary (executable) entry without a complete set of sources?
Entries that do not include a complete set of sources are ineligible for awards.
Is MATLAB available in the test environment?
MATLAB will be available when the scoring interface enviroment re-opens.
A working example of a Challenge entry written in m-code is available for study and as a starting point for developing your own m-code entry. The m-code in the sample works using either MATLAB or Octave.
Will your test environment run multi-threaded code on multiple cores? If so, how many cores will your test environment have and how much memory? Will pthread libraries be pre-installed?
See the details about the test environment here. Briefly, the VMs have dual-core CPUs with 2GB of (shared) RAM. The complete list of pre-installed packages includes libpth20.
Will your linux test box have Qt installed, and if so what version?
No. Your entry should not require a GUI since it will run non-interactively under the control of your setup.sh and next.sh scripts.
Will the wfdb header files and libraries be pre-installed on the test machines such that we can just #include <wfdb/wfdb.h> in C code and link with -lwfdb?
Yes.
Across the training samples there are 46 non-normal beats, variously annotated as "V", "S", and "Q". Are we supposed to try to detect and annotate aberrant beats? Would the prediction error be calculated the same if we annotate them as normal, "N"?
Yes, and yes. For this Challenge, it is not necessary to distinguish between different types of beats, but you should try to detect all of the beats including any abnormal beats.
The annotated locations of the QRS complexes appear to be consistently about midway between the peak at R and the trough at S. Is there a reason the annotations are not located at the point closest to the maximum R amplitude? Is there a formal definition for the location of the QRS complex that you can specify?
Most of the reference annotations were initially generated using a QRS detector that placed them at the centroid of a lag-corrected filtered version of the ECG signal. If the complexes are monophasic, these annotations will appear at or very near the major extremum; otherwise, they will appear at intermediate locations. This approach was chosen for stability with respect to observed fluctuations of the mean cardiac electrical axis that occur as a result of physical motion of the sensing electrodes relative to the heart, such as during normal respiration.
All of the reference annotations were visually reviewed, and corrected where necessary. Some of the initial annotations were repositioned, some were removed, and additional annotations were added (especially when necessary because of severe noise in, or loss of, the ECG) as a result of these reviews. The expert annotators were able to place their annotations as they wished.
It should be emphasized that a correct detection of a beat by a challenge entry does not require determining the precise location of any specific feature; it is sufficient for your software to place its beat annotation anywhere within 150 ms of a reference beat annotation, which will be placed within the observed or expected QRS complex in the ECG signal.
Is there some kind of listserv for participants to post questions and comments?
We hope to launch project-specific wikis on PhysioNetWorks within a few weeks, and the Challenge project will be one of the first to make use of this feature. Until then, please send your questions by email, and if they are of general interest to participants, we will post them (anonymously) and their answers here.
May I use code or complete applications (such as gqrs or wabp) from the WFDB Software Package in my entry?
Yes. You are not required to reinvent anything that you can find in existing open-source software. (The sample entry is gqrs, so you will need to add something more in order to improve on it!)
All of the software that has been preinstalled in the test environment is open-source software, and you may use any of it in your entry.
Although the Challenge rules do not impose any restrictions on the inclusion of existing open-source code in your entry, please be careful to abide by the original author's license. If the license appears to restrict how the code may be used, it may not be an open-source license. Be sure to acknowledge the authors of any code you incorporate in your entry, and cite the original sources in any publications that describe or make use of your entry.
Is a list of names of signals in the hidden test set available?
We are not currently planning to provide such a list, but the first signal in each record will be an ECG signal, and will usually be marked as "ECG". The non-ECG signals will include a variety of pulsatile signals, often including "ABP" (radial arterial blood pressure) and "PLETH" or "PPG" (fingertip photoplethysmogram). Your code may be able to recognize other signals that also contain (quasi-)periodic waveforms that have consistent relationships with the ECG. There may be other signals such as "RESP" (respiration) that are less clearly coupled with cardiac activity. We are hoping to attract entries that discover relationships among the signals and exploit those relationships where appropriate, even in situations in which the signals may be mislabeled (yes, this is also possible -- it's the real world!).
Can a record contain more than one signal with the same name (for example, two or more "ECG" signals)?
In some cases, there may be multiple signals of the same type, but they will never have identical names (for example, two ECG signals in the same record will usually be labeled as "ECG0" and "ECG1").
What does '.../fstat.m shadows a core library function' mean in my entry's diagnostic output'?
This message appears when running an m-code entry, and it results from the presence of the fstat function in Octave's statistics package, which hides a deprecated fstat function that has not yet been removed from Octave's core library. The warning is harmless and will not prevent your entry from running; if your entry does not run successfully, look for another cause.
In the very unlikely case that you wanted to use the deprecated fstat, note that Octave's stat function now accepts either a filename (as always) or a file descriptor (as the deprecated fstat did previously).
Is something wrong with training set record 116?
This record is 1 minute and 49 seconds long, unlike the others in the training set, almost all of which are a full 10 minutes in length. Your entry should be able to handle occasional short records, which may also occur in the test set.
How can I manually install an Octave package in the test environment?
Remember that the test environment does not include a network interface. You must submit any code that your entry needs as part of your entry.tar.gz or entry.zip file, since your entry will not be able to download anything at run time. Follow these steps:
- Download the required package tarball(s) from Octave-Forge and add them to your entry's main directory (the same directory that contains setup.sh).
- Edit your setup.sh script, adding these commands:
STR="${OCTAVE} \"pkg prefix ~ ~;pkg local_list ~/.octave_packages;pkg install YOUR_PACKAGE.tar.gz;quit;\"" echo "$STR" # optional, but helpful for diagnosing problems if they occur eval ${STR}
Any packages you install will remain in the test environment only until your test run has ended; if you submit another entry later, you must reinstall your packages if you still need them.
This FAQ was contributed by Christoph Hoog Antink. The packages that are known to have been installed sucessfully using this method with Octave 3.6.2 during Phase I were:
- general-1.3.2.tar.gz
- specfun-1.1.0.tar.gz
- control-2.4.2.tar.gz
- signal-1.2.2.tar.gz
Note that these four packages are now preinstalled. If your Phase I entry installed any of them, or any other preinstalled package, you will need to remove the code that does so in Phases II and III, since attempting to reinstall an installed package will result in an error, reported in your diagnostc output like this:
error: load_packages: A(I) = X: X must have the same size as I error: called from: error: /usr/share/octave/3.6.2/m/pkg/pkg.m at line 2142, column 21 error: /usr/share/octave/3.6.2/m/pkg/pkg.m at line 401, column 7
Is it possible to change the input sampling frequency using setsampfreq in an m-code entry?
The setsampfreq function is part of the WFDB library, and it is intended to be used by applications written in C or other compiled languages that are linked to the WFDB library. You can write your entry in one of these languages and link it to the WFDB library.
The WFDB Toolbox for MATLAB and Octave provides an m-code interface to many such applications. WFDB Toolbox functions allow you to run these applications as separate processes, capturing their outputs in MATLAB/Octave arrays. If the application calls setsampfreq, its output (the contents of the array(s) that it fills) will reflect this; but the effects of calling setsampfreq do not persist beyond the process in which it is executed, and each call to a function in the Toolbox is executed in its own process — so this will probably not give you the results you may be hoping for.
As noted above, you can either write your code so that it checks the input sampling frequency and properly handles inputs of different sampling frequencies (which is the preferred approach), or you can resample to your desired sampling frequency. If you choose to resample, we don't recommend trying to do this in m-code (it's computationally intensive, and it will be too slow). Rather, write your next.sh script so that it runs the WFDB application xform (which is preinstalled in the test environment) to create a new record resampled at the frequency you want, before starting Octave; then analyze the resampled record with Octave. If you take this approach, remember that the annotation file that your entry generates must include timestamps in the units of the original record's sampling intervals, so your entry will need to compensate for the effects of resampling.
In which folder (directory) are the .hea and .dat input files? Should the .qrs output files be written to this folder?
Your entry is unpacked into your home directory in the test environment, so your home directory contains your setup.sh and next.sh scripts. When these scripts are started, the current directory is your home directory. The .dat and .hea files are copied into your home directory after your setup.sh script exits. Your next.sh script must write the .qrs files into the same directory.
How do you handle entries that contain proprietary (non-free) code?
Our original plan was simply to treat any such entries as invalid. This plan has been modified slightly for Phase I only.
It is not possible for the autoscorer to identify non-free code automatically, and it is not feasible for us to review entries as they arrive to determine if they contain non-free code. One purpose of the hiatus between Phases I and II was to allow us time to review the top-scoring entries for the presence of non-free code before announcing the official rankings. We did not expect to discover any non-free code, but we found that several participants had submitted entries containing MathWorks proprietary code, and further investigation uncovered other sources with copyright statements that did not indicate the authors' permission to redistribute their code.
We don't wish to revoke the provisional qualification of the authors of these entries as official participants, especially since it appears that inclusion of non-free code may have been inadvertent in some or all cases -- so we have allowed these entries, if they were scored, to meet the eligibility requirement for submitting a scorable Phase I entry.
Nevertheless, we don't wish to give awards for entries that do not comply with the rules; this would be unfair to the majority of participants who were careful to follow the rules. Therefore, no awards and no official rankings will be given for entries containing non-free code (whether or not the code was used in the entry).
If I use up all of my entries, can I register again using another email address and get more entries?
Please don't do this! The limit on the number of entries:
- is necessary in order to make it feasible to evaluate entries within a reasonable time as deadlines approach,
- reduces the opportunities for overfitting and consequent development of non-generalizable solutions, and
- is an important part of establishing a fair and level playing field for the Challenge.
Obviously we cannot prevent use of multiple email addresses to circumvent the limit on entries, but we consider such behavior abusive and disrespectful to other participants. In Phase I, we have disqualified excess entries. If we discover that this has occurred in Phases II or III, we will disqualify offending participants.
Note, however, that participants are allowed (and encouraged) to join forces to combine the best features of complementary approaches. If you wish to collaborate with another participant or team of participants, each of the team members may use his or her remaining entries jointly with other team members. Collaborators must be particularly careful to list all team members in their AUTHORS files to avoid disqualification.
Why are my Phase II scores so much worse than my Phase I scores, for the same code?
The Challenge is becoming even more challenging! The Phase II test set consists of 200 records: the 100 records in the Phase I set, and 100 additional records, many of which are considerably more difficult than the average among the first 100. Everyone's scores are likely to be significantly lower in Phase II. This will happen again in Phase III, when another 100 records are added to the test set.
Why do not I get 'output does not match' errors ?
When I submit my entry through PhysioNetWorks I get an error message similar to:
Challenge 2014 entry running ...
next.sh 100: output does not match submission
octave --quiet --eval "challenge('100',0); quit;" 2>&1
This error message indicates that, for whatever reason, the annotations files created by running your entry in our Sandbox environment do not match annotation files submitted with your entry. Please make sure that your submission runs properly in the Sandbox bootable ISO.
We strongly recommend that you use this Sandbox environment to generate the annotation files that you submit along with your entry. You can use a virtual environment, such as VirtualBox, to have the ISO quickly installed and running as a guest on your machine.
Why does my entry seems to get hanged, printing a 'save to `octave-core' complete' error ?
My entry never finishes running and I get a message similar to:
core library function
warning: time stamp for `/home/vmuser/challenge.m' is in the future
attempting to save variables to `octave-core'...
save to `octave-core' completeChallenge 2014 entry running ...
next.sh 100: output does not match submission
octave --quiet --eval "challenge('100',0); quit;" 2>&1
You entry has timed out (the results will not be reported). The reason you are seeing this error message is because Octave has received an interrupt signal from the VM and is attempting to save the current state of the workspace into a file. Currently, this prevents our monitoring software from proceeding with the standard 'time-out' message. In order to avoid this behaviour, please add the following lines of code to the top of your 'challenge.m' file:
[~,config]=wfdloadlib; %You may have this line already if(config.inOctave) crash_dumps_octave_core(0); end
Where can I find more training data to test my algorithm ?
The datasets from Phase I and Phase II are a subset of the dataset of Phase III. Therefore, it is not possible to expand the training records by opening the Phase I and II datasets. You can use any data available at PhysioNet to expand your training records, the pbsearch tool should help with quickly identifying records of interest.
How are the records and header files loaded in the testing environment ?
The *.dat and *.hea files used for generating the scoring on the VM are loaded into the top directory of the unzipped entry location. These files are *not* loaded into the './challenge/2014/set-p/' directory (where the user submitted annotation files reside). Therefore, to read the data files correctly, please make sure your entry has either just the record name, or the full path to the record. For example, reading a data file from Octave, either of these methods should work (assuming challenge.m resides on the top directory of your unzipped entry file):
%Method 1: recordName='100'; [tm,sig]=rdsamp(recordName); %Method 2: Using full path to record file recordName='100'; recordPath=[pwd '/']; [tm,sig]=rdsamp([recordPath recordName]);
How can I quickly load WFDB record data into Octave ?
Is there an alternative to RDSAMP.m for quickly loading signal data into Octave ? When I try to use RDSAMP, Octave takes a few seconds to load individual data into the workspace, is there a faster way?
You can use the WFDB2MAT command in order to convert the record data into a *.mat file, which can then be loaded into Octave's workspace using the LOAD command. This sequence of procedures is quicker (by several orders of magnitude) than calling RDSAMP. For example, assuming that WFDB2MAT is properly installed on your system (like in the Sandbox bootable ISO), and that the record files 100.dat and 100.hea are in the current directory:
%Generate the 100m.mat and 100m.hea fiels from the *.dat and *.hea files
system('wfdb2mat -r 100 -H');
%Load the new file into Octave's workspace. This will create a 'val' variable containing the data
load('100m');
Note that this will load the signal data in raw units. To convert from raw units to physical units you will need to follow the instructions described when running this command (ie, subtract the given 'base' of a signal and divide by the given 'gain' of that signal).
What were the changes to the scoring system?
The scoring interface to the 2014 PhysioNet Challenge has been re-opened for scoring on January 23, 2015. If you entry takes more than 12 hours to score please contact us. Please note carefully the following editions:
- The original training set has been augmented with 100 records from the original test set. Thus a new test, of 200 records, has been created that does not include these newly released records (the original test set had 300 records).
- Your scores will be calculated on this new test set of 200 records. When reporting your scores for the special issue on Physiological Measurement, please use the scores obtained from the new test set. In most cases, they will be higher than those generated based on the old test set.
- There will be a limit of 20 entries that can be scored between now and February 27, 2015. After this date, we will change this limit to a small fixed number of entries per month.
- Entries were previously limited to 40 seconds of runtime per record and total time of about 3 hours. This has been changed. Entries are now limited to 6*10^10 CPU instructions per record and the total time limit has been removed. Entries that were able to complete within 40 seconds should finish within this limit, feedback statistics on the number on the number of CPU instructions used by your entry is provided via the web interface.
Challenge Results
The top results of the follow-up entries as well as all entries from phases I-III (at the end of February 2015) were achieved by Urska Pangerc (93.64), Alistair Johnson (91.50), Sachin Vernekar (90.97), Christoph Hoog Antink (90.70), and Abid Rahman (90.16). The complete updated rankings are now available including follow-up entries as well as all entries from phases I-III.
Papers
The following paper is an introduction to the challenge topic, with a summary of the challenge results and a discussion of their implications. Please cite this publication when referencing the Challenge.
Robust Detection of Heart Beats in Multimodal Data: The PhysioNet/Computing in Cardiology Challenge 2014
George Moody, Benjamin Moody, Ikaro Silva
Papers presented at Computers in Cardiology 2014 have been made available by their authors under the terms of the Creative Commons Attribution License 3.0 (CCAL). See this page for a list of the papers. We wish to thank all of the authors for their contributions.
Access
Access Policy:
Anyone can access the files, as long as they conform to the terms of the specified license.
License (for files):
Open Data Commons Attribution License v1.0
Discovery
Topics:
multiparameter
challenge
ecg
Corresponding Author
Files
Total uncompressed size: 0 B.
Access the files
- Download the ZIP file (506.3 MB)
- Access the files using the Google Cloud Storage Browser here. Login with a Google account is required.
-
Access the data using the Google Cloud command line tools (please refer to the gsutil
documentation for guidance):
gsutil -m -u YOUR_PROJECT_ID cp -r gs://challenge-2014-1.0.0.physionet.org DESTINATION
-
Download the files using your terminal:
wget -r -N -c -np https://physionet.org/files/challenge-2014/1.0.0/
-
Download the files using AWS command line tools:
aws s3 sync s3://physionet-open/challenge-2014/1.0.0/ DESTINATION
| Name | Size | Modified |
|---|---|---|
| octave | ||
| papers | ||
| set-p | ||
| set-p2 | ||
| sources | ||
| entry.tar.gz (download) | 156.6 KB | 2019-04-17 |
| entry.zip (download) | 173.5 KB | 2019-04-17 |
| example-search.txt (download) | 9.4 KB | 2019-04-17 |
| figure1.png (download) | 47.2 KB | 2019-04-17 |
| octave-sample-entry.zip (download) | 120.7 MB | 2019-04-17 |
| package-list.txt (download) | 20.0 KB | 2019-04-17 |
| rankings-20150413 (download) | 2.6 KB | 2019-04-17 |
| set-p-atr.tar.gz (download) | 74.8 KB | 2014-01-07 |
| set-p.tar.gz (download) | 111.3 MB | 2013-12-03 |
| set-p.zip (download) | 111.3 MB | 2013-12-03 |
| talk-instructions.txt (download) | 1.7 KB | 2019-04-17 |
| training.zip (download) | 112.7 MB | 2019-04-17 |